Download
The ScoresDecomposer.py script summarizes the scoring components from Dock scoring files, OEDocking scoring tables and Gold log files into a table-like file to be used as input to CompScore. This script is freely available for download from here. System requirements Before running ScoresDecomposer be sure that your system meets the following requirements:
- Python >=3.6 (Conda >= 4.4.10 recommended)
- NumPy >= 1.14.0
Supported scoring functions
Currently, ScoresDecomposer.py supports the extraction of scores components from the scoring functions listed in the below table. The components of these scoring functions are extracted according to files formats and identifying keywords.
| Docking program | Scoring functions |
| Dock 6.8 | Grid, Contact, Continuous, Hawkins, PBSA, SASA |
| Gold | PLP, GoldScore, ChemScore, ASP |
| OEDocking | Shapegauss, ChemScore, ChemGauss 3, ChemGauss 4, PLP |
Running
The script is run as:
python ./ScoresDecomposer.py -i <Input File> -o <Output Scores Table>
The OutputFile will contain a text file that can be used as Data file for the CompScore Web Service. The format of the input file for “ScoresDecomposer.py” is described in teh next section.
Input file
The input file contains information regarding the output files from docking calculations. This information includes the file location, the docking program that produced the file and the scoring function the file corresponds to. The file contains information of one scoring file per line with fields separated by tab. Columns is the input file are organized as follow:
- Column 1: Location of the scoring file relative to current directory
- Column 2: Docking program ID. The IDs of the docking programs are Dock35 for Dock 3.6, Dock6 for Dock 6.8, OpenEye for OEDocking and GOLD for Gold. The use of these IDs are mandatory.
- Column 3: Scoring function ID. Each scoring function from the above listed programs are identified as:
| Param | Scoring function | Scoring function ID |
| Dock35 | Dock 3.6 | Dock35 |
| Dock6 | Grid | D6Grid |
| Dock6 | Contact | D6Contact |
| Dock6 | Continuous | D6Continuous |
| Dock6 | Hawkins | D5Hawkins |
| Dock6 | PBSA | D6Pbsa |
| Dock6 | SASA | D6SASA |
| OpenEye | ChemGauss 3 | OEChemgauss3 |
| OpenEye | ChemGauss 4 | OEChemgauss4 |
| OpenEye | ChemScore | OEChemScore |
| OpenEye | PLP | OEPLP |
| OpenEye | Shapegauss | OEShapegauss |
| GOLD | ASP | GASP |
| GOLD | ChemScore | GChemscore |
| GOLD | GoldScore | GGoldscore |
| GOLD | PLP | GPlp |
It is important that IDs for programs and scoring functions are correctly specified in the input file, otherwise the ScoresDecomposer.py will fail to generate a scores components summary file. A typical input file for this script will look like:
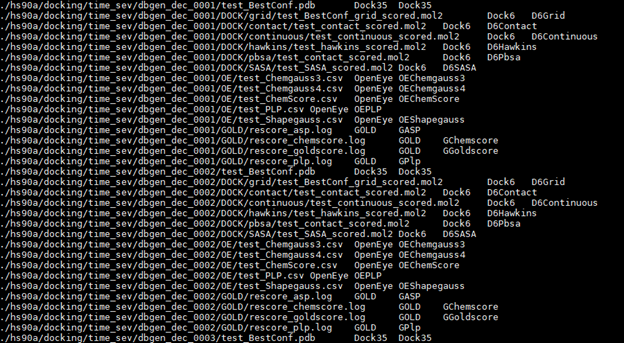
Output file
The OutputFile is a text file that can be used as Data file for the CompScore Web Service. This file is organized as:
- First row: data heading, i.e scores components IDs.
- Column 1: The ID of each compound.
- Column 2: The classification of the compounds in either ligand (1) or decoy (0).
- Column 3: The value of the number of heavy atoms.
- Column 4: to last: Scoring components values.
Dock 6.8 Grid scoring files are used as the reference files for the extraction of the classification and number of heavy atoms of the compounds. Compounds not present in a Dock 6.8 Grid scoring file will be assigned a number of 999999. Given that all calculations for CompScore validation were performed with the DUD-E database, for classification extraction if the “dbgen_dec” pattern is found in the reference file the compounds in that file are classified as decoys (0). On the other hand, if the “dbgen_lig” pattern is found in the reference file the compounds in that file are classified as ligands (1).
Compounds not present in a Dock 6.8 Grid scoring file or those present but for which no “dbgen_dec” or “dbgen_lig” patterns are found in their paths a classification of -1 is assigned. In cases where some compounds can’t be assigned to either the ligands or decoys groups and the OutputFile of ScoresDecomposer.py are required for a CompScore GA optimization run, the user can manually add the compounds classification latter. Finally, the IDs that ScoresDecomposer.py assign to each docking score component are listed here .
